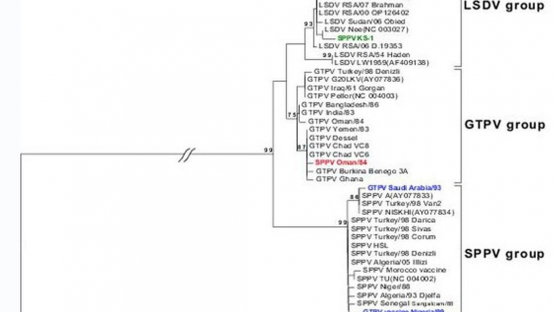
The data revealed that the genetic information within each of these two genes was sufficient to produce phylogenetic trees with similar topologies to the one obtained with the full genome information of CaPVs. For the full genome tree, three clusters composed of LSDV, GTPV and SPPV were found (see example of the RPO30 gene in Figure 2). This suggests that by sequencing each of these genes for a given CaPV strain, it is possible to determine its genotype and assign it to one of the three CaPV groups, using a phylogenetic reconstruction. Given the large number of freely available bioinformatics programmes, (some are web-based) and the development of high throughput sequencing methods, it is feasible for some laboratories in developing IAEA MS to use this approach to better characterize CaPVs. For example, these laboratories could use the primers sets that have been designed by APHL to amplify and sequence one or both of these two genes to characterize virus isolates from disease outbreaks in their own region. Subsequently, the sequencing data could then be used together with the set of CaPV sequences that have been deposited in GenBank by APHL scientists and others to generate phylogenetic trees using freely available software to classify their isolates.
Another important finding was the presence of viral isolates that were located outside the group corresponding to their host of origin. While this provides evidence for cross-infection with CaPVs such as SPPV infecting goats or GTPV infecting sheep, it also shows the weakness of the system for naming and classifying CaPVs that is based solely on the host species from which the virus was first isolated. The results of this collaborative work strengthen the need to use molecular-based tools for CaPVs strain designation and classification. This work has also shown the potential of the species-specific signatures in both of these genes to design simple CaPVs genotyping tools to screen large number of samples during outbreaks without further gene sequencing. The most interesting signatures were the exclusive presence of a 21-nucleotide deletion on the RPO30 gene on all SPPV group members, which can be used to differentiate them from LSDV and GTPV, and the presence of species-specific signatures on the CaPVs GPCR, which represent targets for genotyping with hybridisation probes.
These results have been disseminated to the scientific community through two peer review publications (Journal of General Virology, 2009, 90, 1967-1977 and Veterinary Microbiology (2010, doi:10.1016/j.vetmic.2010.09.038).
The identification of these two genes as potential genotyping targets for CaPVs represents a significant advance in the biology of CaPVs, contribution that will facilitate understanding their epidemiology and assist in rapid decision-making in identifying a suitable vaccine for more efficient control. Certain species-specific signatures that were found within each of these genes were used by the APHL to design simple and cost effective molecular based procedures for CaPVs strain identification, which will not require any sequencing work.